Overview
Our laboratory uses NMR spectroscopy and related biophysical methods to characterize the structure and dynamics of proteins involved in signal transduction and the regulation of gene expression and in the synthesis/degradation of carbohydrates. We receive our funding through the Canadian Cancer Society, the Natural Sciences and Engineering Research Council of Canada (NSERC), and the Canadian Institutes of Health Research (CIHR).
Structural Bases of Ets-1 Regulation: Auto-Inhibition, Protein Partnerships and Post-translational Modifications.
The Ets family of transcription factors and oncoproteins has emerged as a paradigm for the study of the gene expression in eukaryotes. The activities of these proteins are dependent upon numerous possible protein-protein and protein-DNA interactions, thus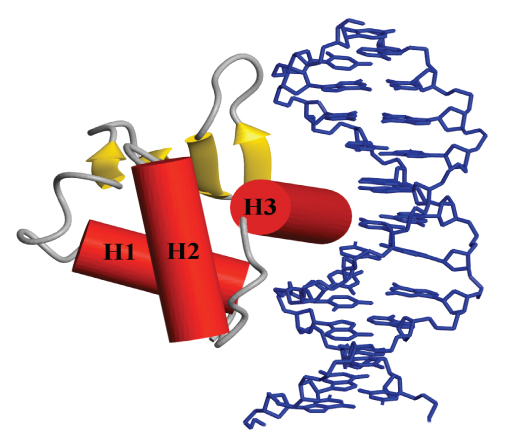 providing an ideal system for investigating structurally the mechanisms of transcriptional regulation and its relationship to oncogenesis. In particular, we are focusing on the control of murine Ets-1 through auto-inhibition of DNA binding, protein partnerships, and post-translational modifications, including phosphorylation and sumoylation, via signal transduction cascades.
providing an ideal system for investigating structurally the mechanisms of transcriptional regulation and its relationship to oncogenesis. In particular, we are focusing on the control of murine Ets-1 through auto-inhibition of DNA binding, protein partnerships, and post-translational modifications, including phosphorylation and sumoylation, via signal transduction cascades.
Most significantly, we have developed a model for the allosteric regulation of Ets-1 in which increasing multi-site phosphorylation of a dynamic, unstructured serine rich region leads to a progressive decrease in promoter affinity due to transient interactions that stabilize a labile auto-inhibitory helix appended to the DNA binding ETS domain. This defines a “rheostatic” (rather than off/on) mechanism for cell signaling by fine tuning transcription at the level of DNA binding.
In parallel, we have also undertaken detailed structural and dynamic analyses of the PNT domain from several Ets transcription factors. The Ets-1 PNT domain serves as a docking motif for the MAP kinase ERK2, thereby facilitating phosphorylation of two adjacent phosphoacceptors. These modifications shift a conformational equilibrium of a dynamic helix in the PNT domain from a more closed to a more open state, thereby facilitating binding to the TAZ1 domain of the co-activator CBP. This phospho-switch mechanism also highlights the role of appended dynamic elements to conserved structural domains in the evolution of signalling pathways.
Enzymatic Mechanisms of Glycoside Hydrolases and Glycosyl transferases.
Carbohydrates are essential to life and thus the enzymes that catalyze the breakdown (glycoside hydrolases) and synthesis (glycosyl transferases) of these molecules play critical roles in biology and have many practical applications in medicine and biotechnology.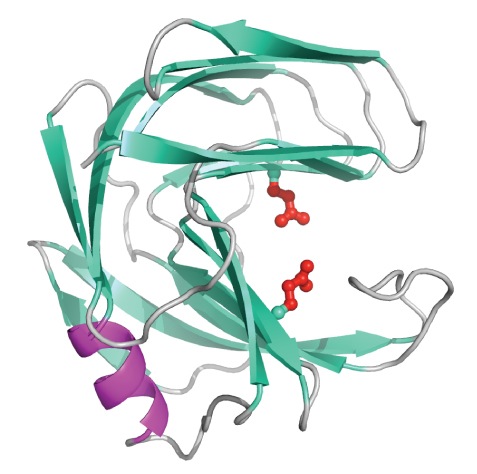 For example, cellulose and related xylan are the most abundant forms of carbon on earth. Accordingly, the hydrolases that breakdown these carbohydrates have enormous “green chemistry” roles in food, textile, paper, and fuel production. Conversely, compounds that inhibit the action of glycoside hydrolases and glycosyl transferases may help control diseases such as microbial infections, diabetes, and cancers. Therefore, these enzymes remain the subject of intensive research directed towards understanding and manipulating the mechanisms by which they build or degrade their carbohydrate substrates.
For example, cellulose and related xylan are the most abundant forms of carbon on earth. Accordingly, the hydrolases that breakdown these carbohydrates have enormous “green chemistry” roles in food, textile, paper, and fuel production. Conversely, compounds that inhibit the action of glycoside hydrolases and glycosyl transferases may help control diseases such as microbial infections, diabetes, and cancers. Therefore, these enzymes remain the subject of intensive research directed towards understanding and manipulating the mechanisms by which they build or degrade their carbohydrate substrates.
The general theme of our research program involves using NMR spectroscopy as a “molecular microscope” to examine several model glycoside hydrolases and glycosyl transferases. Our primary goals are: (i) to define the electrostatic changes occurring along their reaction pathways by measuring the pH-dependent charge states (pKa values) of catalytic amino acids; (ii) to investigate the roles of molecular motions in substrate binding and subsequent breakdown or synthesis; and (iii) to use this information to tailor these enzymes for biotechnological applications such as paper bleaching under high pH conditions.
Biomolecular NMR spectroscopy.
Our research group has dedicated 500, 600, and (soon to be purchased) 800 MHz instruments fully equipped for all aspects of modern biological NMR spectroscopy.



